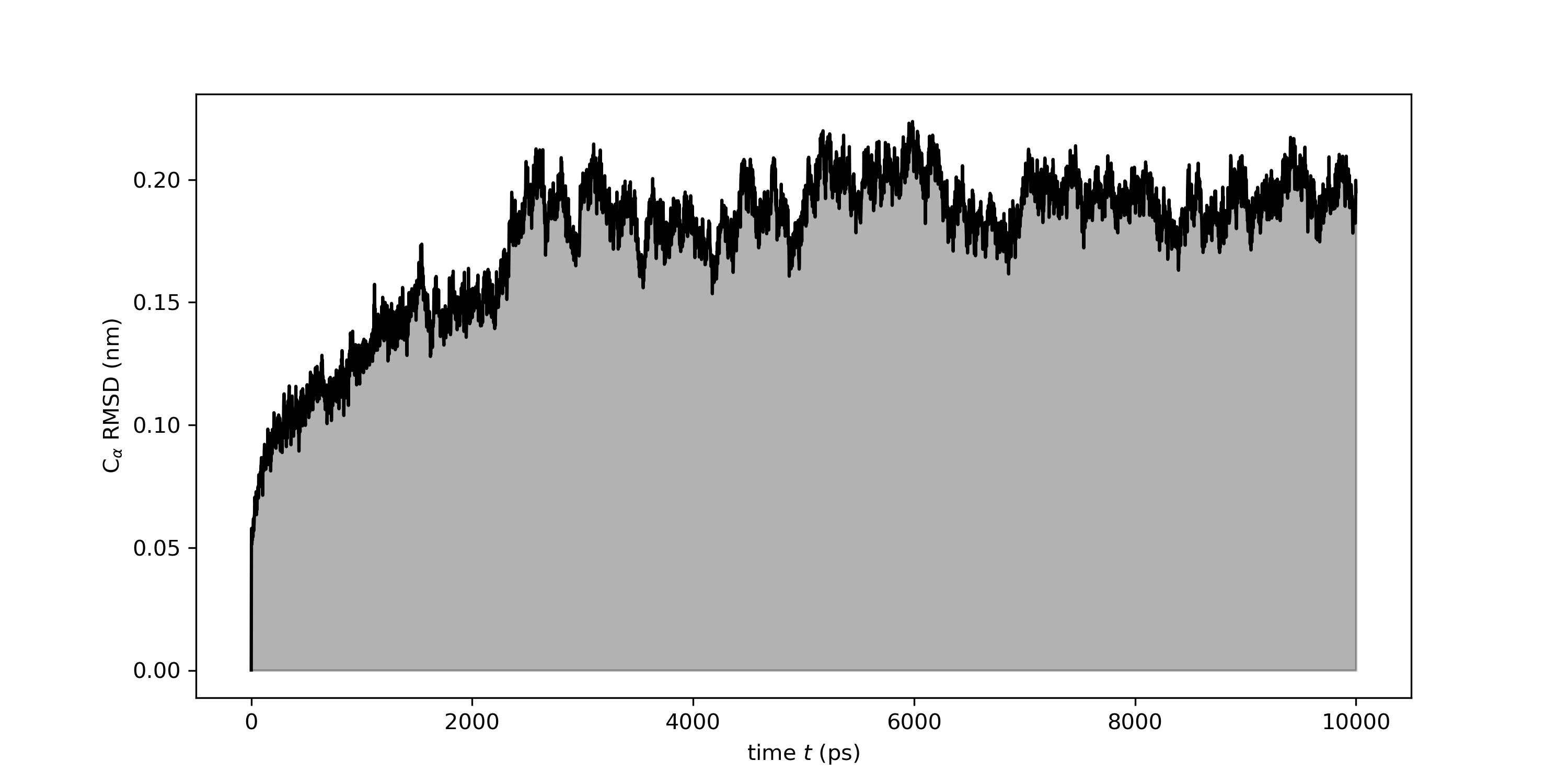
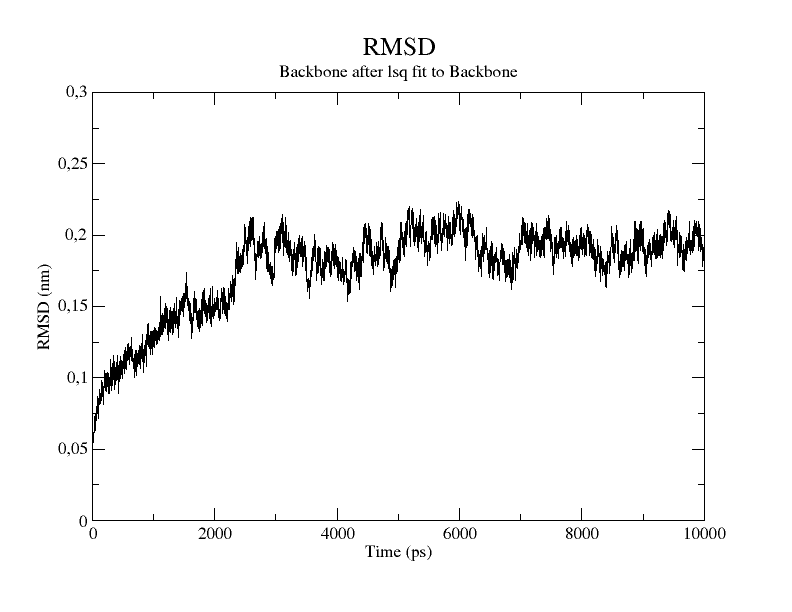
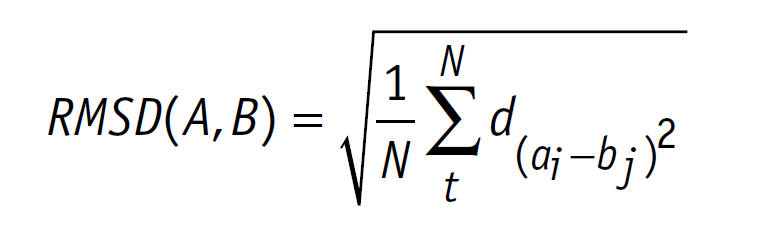Average all-atom root-mean-square deviation (RMSD) calculated over time... | Download Scientific Diagram
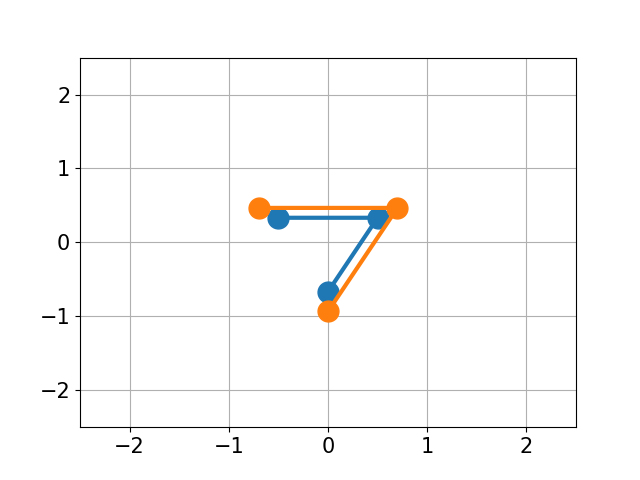
GitHub - charnley/rmsd: Calculate Root-mean-square deviation (RMSD) of two molecules, using rotation, in xyz or pdb format
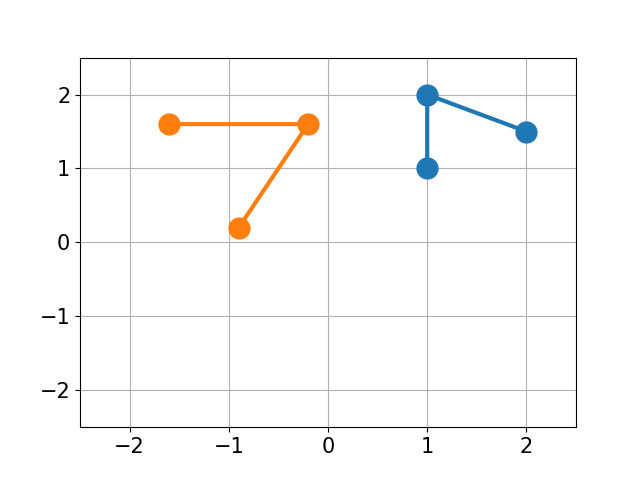
GitHub - charnley/rmsd: Calculate Root-mean-square deviation (RMSD) of two molecules, using rotation, in xyz or pdb format
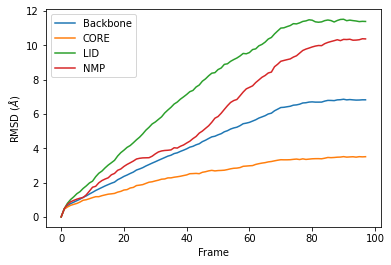
Calculating the root mean square deviation of atomic structures — MDAnalysis User Guide documentation



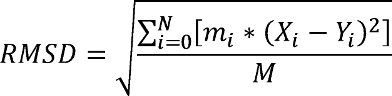



![PDF] A Normalized Weighted RMSD for Measuring Protein Structure Superposition | Semantic Scholar PDF] A Normalized Weighted RMSD for Measuring Protein Structure Superposition | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/efd54bbf13038ecf12dfdabc54a9267afffa3598/4-Figure1-1.png)